
ΑΙhub.org
Engineering molecular interactions with machine learning
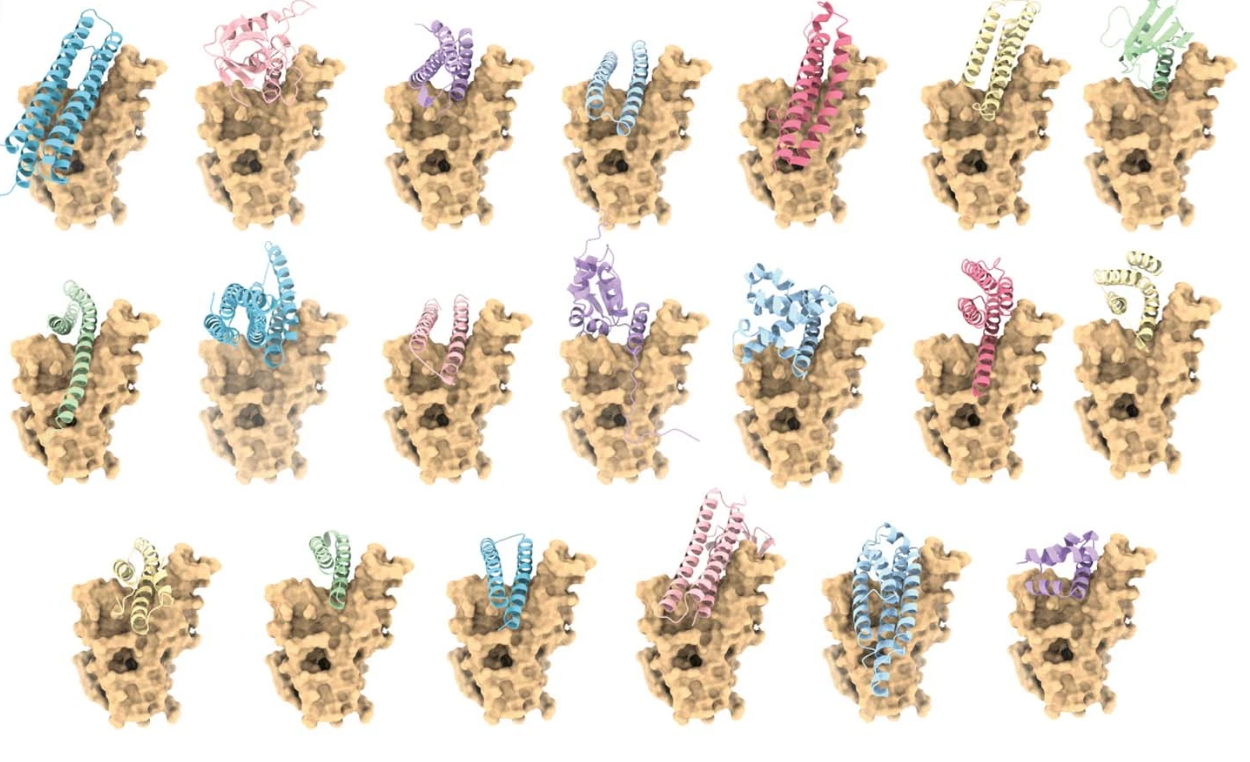 Receptor-binding domain-binder designs displayed on yeast. From De novo design of protein interactions with learned surface fingerprints. Reproduced under a CC BY 4.0 licence.
Receptor-binding domain-binder designs displayed on yeast. From De novo design of protein interactions with learned surface fingerprints. Reproduced under a CC BY 4.0 licence.
By Celia Luterbacher
In 2019, scientists in the joint School of Engineering and School of Life Sciences Laboratory of Protein Design and Immunoengineering (LPDI) led by Bruno Correia developed MaSIF: a machine learning-driven method for scanning millions of protein surfaces within minutes to analyze their structure and functional properties. The researchers’ ultimate goal was to computationally design protein interactions by finding optimal matches between molecules based on their surface chemical and geometric “fingerprints”.
Four years later, they have achieved just that. In a paper published in Nature, they report that they have created brand-new proteins called binders that are designed to interact with four therapeutically relevant protein targets, including the SARS-CoV-2 spike protein.
Engineering a perfect molecular match
Physical interactions between proteins influence anything from cell signalling and growth to immune responses, so the ability to control protein-protein interactions is of great interest to the fields of biology and biotechnology. While textbook depictions of protein binding may look as simple as fitting puzzle pieces together, the reality is more complex: protein surfaces vary widely and are dynamic, making it hard to predict how and where binding events will occur.
“A puzzle piece is two-dimensional, but with protein surfaces, we are looking at multiple dimensions: chemical composition, such as positive versus negative charge interactions; shape complementarity, curvature, etc.,” explains LPDI PhD student and co-author Anthony Marchand.
“The idea that everything in nature that binds is complementary – for example, a positive charge binds with a negative charge – has been a long-standing idea in the field, which we captured in our computational framework.”
To design novel protein binders, the researchers used MaSIF to create protein surface “fingerprints”, and then identified complementary surfaces for key protein target sites from a database of fragments. They then digitally grafted the fragments onto larger protein scaffolds, and selected the resulting binders predicted to interact best with their targets. After synthesizing and testing these selected binders in the lab, the researchers were able to confirm the computationally generated hypothesis.
“The fact that we’re able to design novel, site-specific protein binders in just a couple of months makes this method very interesting for therapeutics. It’s is not just a tool: it’s a pipeline,” Marchand says.
“Straight from the computer”
The researchers were developing protein binders for three major cancer immunotherapy targets when the COVID pandemic hit, so they added the SARS-CoV-2 spike protein to their list. Using their approach, the four binders they produced displayed excellent affinities for their targets.
MaSIF’s success rate, combined with its speed and ability to produce high-quality, site-specific designs, all demonstrate its therapeutic potential. For example, the ability to generate accurate protein binders so rapidly could be a big advantage for epidemiological applications, as in the case of the SARS-CoV-2 spike protein. Marchand also sees potential for the pipeline to facilitate the development of chimeric antigen receptor (CAR-T) proteins, which can be engineered to allow patient immune cells to target cancer cells.
“Further advances in machine learning methods will help improve our method, but our work today already provides a strategy for developing innovative therapies to benefit patients through the rapid design of protein-based therapeutics – straight from the computer.”
Read the research in full
De novo design of protein interactions with learned surface fingerprints, Pablo Gainza, Sarah Wehrle, Alexandra Van Hall-Beauvais, Anthony Marchand, Andreas Scheck, Zander Harteveld, Stephen Buckley, Dongchun Ni, Shuguang Tan, Freyr Sverrisson, Casper Goverde, Priscilla Turelli, Charlène Raclot, Alexandra Teslenko, Martin Pacesa, Stéphane Rosset, Sandrine Georgeon, Jane Marsden, Aaron Petruzzella, Kefang Liu, Zepeng Xu, Yan Chai, Pu Han, George F. Gao, Elisa Oricchio, Beat Fierz, Didier Trono, Henning Stahlberg, Michael Bronstein & Bruno E. Correia, Nature (2023).









