
ΑΙhub.org
Using engineered bacteria and AI to sense and record environmental signals
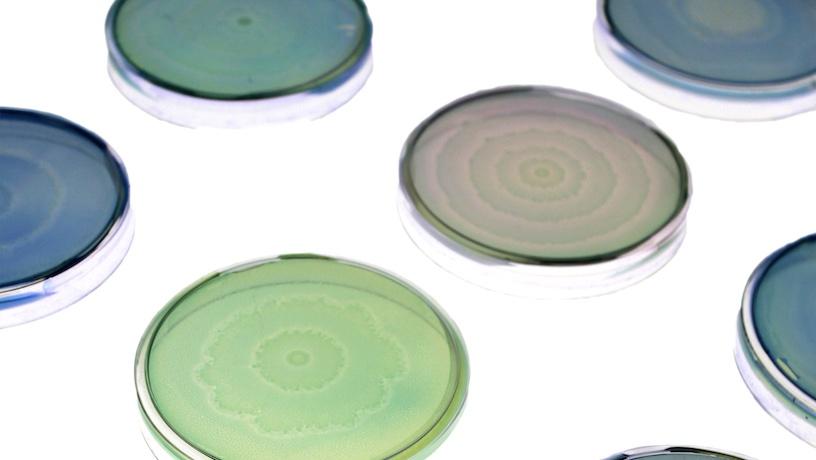 Petri dishes of engineered and native Proteus mirabilis patterns, here stained with colored dyes used for the lab’s bacterial art. Credit: Soonhee Moon, Danino Lab/Columbia Engineering
Petri dishes of engineered and native Proteus mirabilis patterns, here stained with colored dyes used for the lab’s bacterial art. Credit: Soonhee Moon, Danino Lab/Columbia Engineering
By Holly Evarts
Researchers in Biomedical Engineering Professor Tal Danino’s lab were brainstorming several years ago about how they could engineer and apply naturally-pattern-forming bacteria. There are many bacteria species, such as Proteus mirabilis (P. mirabilis), that self-organize into defined patterns on solid surfaces that are visible to the naked eye. These bacteria can sense several stimuli in nature and respond to these cues by “swarming”—a highly coordinated and rapid movement of bacteria powered by their flagella, a long, tail-like structure that causes a whip-like motion to help propel them.
For inspiration, Danino’s team at Columbia Engineering, which has a good deal of experience using synthetic biology methods to manipulate bacteria, discussed where else they might find similar patterns in nature and what their functions might be. They noted how tree rings record tree age and climate history, and that sparked their idea of applying P. mirabilis rings as a recording system. They had also been interested in applying AI to characterize the distinct features of bacterial colony patterns, an approach that they realized could then be used to decode an engineered pattern.
“This seemed to us to be an untapped opportunity to create a natural recording system for specific cues,” said Danino, a member of Columbia’s Data Science Institute (DSI).
In a new study, published May 4 in Nature Chemical Biology, the researchers worked with P. mirabilis, commonly found in the soil and water and occasionally the human gut, known for its bullseye-appearing colony patterns. When the bacteria are grown on a Petri dish of a solid growth media, they alternate between phases of bacterial growth, which make visible dense circles, and bacterial movement, called “swarming” movement, which expands the colony outwards.
Engineering bacteria to sense, respond, and change swarming
Time-lapse movie of the ring pattern formation of the native P. mirabilis strain. The bacterium forms the pattern by alternating between phases of swarming, which expand the colony outwards, and local population growth, which creates dense ring edges. The patterns generally form over 1-3 days and are easily visible to the naked eye. Credit: Daninio Lab/Columbia Engineering.
The team engineered the bacteria by adding what synthetic biologists call “genetic circuits”—systems of genetic parts, logically compiled to make the bacteria behave in a desired way. The engineered bacteria sensed the presence of the researchers’ chosen input—ranging from temperature to sugar molecules to heavy metals such as mercury and copper—and responded by changing their swarming ability, which visibly changed the output pattern.
Using AI to decode swarming pattern
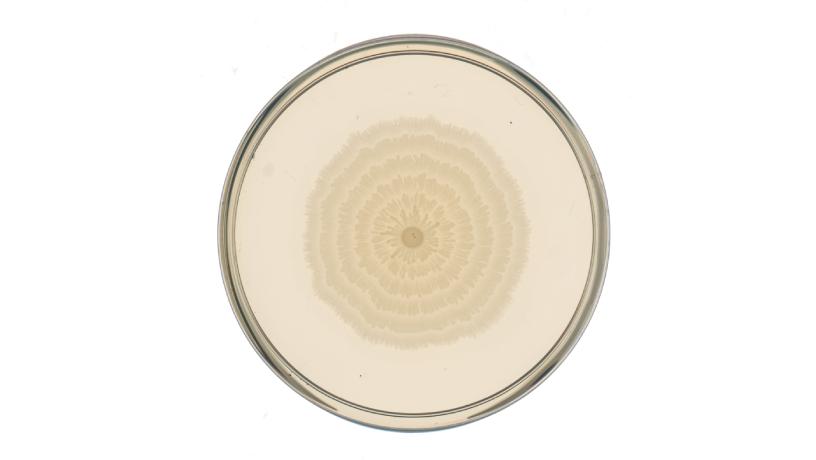 One of the engineered P. mirabilis strains, the “pLac-lrp” strain; when grown on a Petri dish with the molecule IPTG present in the growth medium, as shown here, the strain responds by changing its ring pattern into a pattern of spikes. Credit: Danino Lab/Columbia Engineering.
One of the engineered P. mirabilis strains, the “pLac-lrp” strain; when grown on a Petri dish with the molecule IPTG present in the growth medium, as shown here, the strain responds by changing its ring pattern into a pattern of spikes. Credit: Danino Lab/Columbia Engineering.
Working with Andrew Laine, Percy K. and Vida L. W. Hudson Professor of Biomedical Engineering and a DSI member and Jia Guo, assistant professor of neurobiology (in psychiatry) at the Columbia University Irving Medical Center the researchers then applied deep learning–a state-of-the-art AI technique–to decode the environment from the pattern, in the same way scientists look at the rings in a tree trunk to understand the history of its environment. They used models that can classify patterns holistically to predict, for example, sugar concentration in a sample, and models that can delineate or “segment” edges within a pattern to predict, for example, the number of times the temperature changed while the colony grew.
An advantage of working with P. mirabilis is that, compared to many of the typical engineered bacterial patterns, the native P. mirabilis pattern is visible to the naked eye without costly visualization technology and forms on a durable, easy-to-work-with solid agar medium. These properties increase the potential to apply the system as a sensor readout in a variety of settings. Using deep learning to interpret the patterns can enable researchers to extract information about input molecule concentrations from even complex patterns.
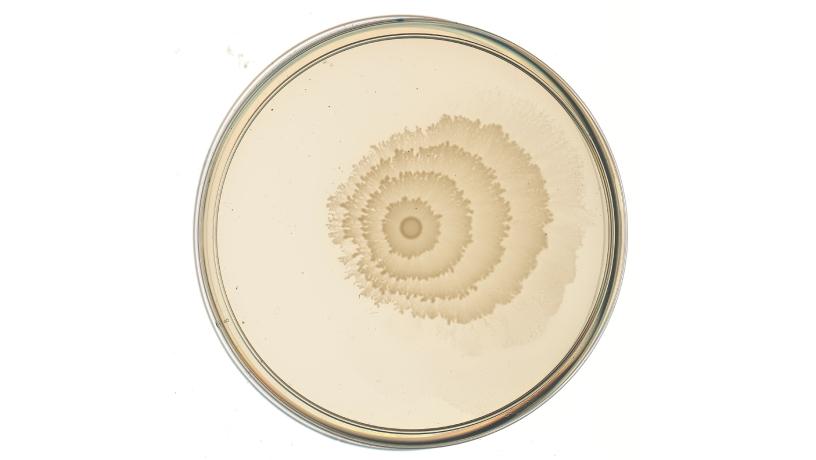 The engineered copper sensing strain, grown with samples of water containing high copper (50 mM) which were added as drops onto the left side of the Petri dish. The copper is sensed by the bacteria, which respond by changing their swarming and thus the ring pattern, such that the invisible presence of copper becomes visible to the naked eye. The pattern on the right side of the plate, with no sample, can be used as a baseline to which the left side’s pattern can be compared. Credit: Danino Lab/Columbia Engineering.
The engineered copper sensing strain, grown with samples of water containing high copper (50 mM) which were added as drops onto the left side of the Petri dish. The copper is sensed by the bacteria, which respond by changing their swarming and thus the ring pattern, such that the invisible presence of copper becomes visible to the naked eye. The pattern on the right side of the plate, with no sample, can be used as a baseline to which the left side’s pattern can be compared. Credit: Danino Lab/Columbia Engineering.
“Our goal is to develop this system as a low-cost detection and recording system for conditions such as pollutants and toxic compounds in the environment,” said Anjali Doshi, the study’s lead author and a recent PhD graduate from Danino’s lab. “To our knowledge, this work is the first study where a naturally pattern-forming bacterial species has been engineered by synthetic biologists to modify its native swarming ability and function as a sensor.”
New approach will advance biotechnology
Such work can help researchers better understand how the native patterns form, and beyond that, can contribute to other areas of biotechnology beyond the area of sensors. Being able to control bacteria as a group rather than as individuals, and control their movement and organization in a colony, could help researchers build living materials at larger scales, and help with the Danino lab’s parallel goal of engineering bacteria to be living “smart” therapeutics, by enabling better control of bacterial behaviors in the body.
This work is a new approach for building macroscale bacterial recorders, expanding the framework for engineering emergent microbial behaviors. The team next plans to build on their system by engineering the bacteria to detect a wider range of pollutants and toxins and moving the system to safe “probiotic” bacteria. Ultimately, they aim to develop a device to apply the recording system outside of the lab.









